Different Model Settings#
The SPyIce package is a software tool that enables 1D finite difference simulation for vertical transport equations. It specifically focuses on thermal diffusion with the influence of salinity and physical properties. The package utilizes the Thomas tridiagonal solver as the solver algorithm. With SPyIce, users can model and analyze the behavior of temperature, salinity, and other relevant variables in a vertical system. It provides a comprehensive framework for studying the thermal diffusion process and its interaction with salinity in various scenarios. Hydra is used to automate the simulation runs of the Sea-Ice Model. It is used to manage and run sea ice simulations, making it easier for users to explore different scenarios and optimize their models.
Import Packages#
[1]:
import os
from pathlib import Path
import matplotlib.pyplot as plt
%matplotlib inline
from omegaconf import OmegaConf
from spyice.utils import create_output_directory
from spyice.postprocess import Analysis, VisualiseModel
from spyice.utils import ConfigSort
from spyice.models import SeaIceModel
from spyice.preprocess import PreProcess
Define Inputs and Project Output paths#
[2]:
# creates a OmegaConf object from a dictionary for fast testing only for parameters: constants, dt, S_IC, iter_max, dz
constants_dict = {
"constants": {"constants": "real"},
"dt": {"dt": 47},
"iter_max": {"iter_max": 10000},
"dz": {"dz": 0.05},
"model": {"is_diffusiononly_equation": False, "is_salinity_equation": True, "is_radiation_equation": True, "is_algae_equation": True, "algae_model_BAL_type": "all"},
"ICBC": {"S_IC": 'S_34', "T_BC": 265.0}
}
config_raw = OmegaConf.create(constants_dict)
config = ConfigSort.getconfig_dataclass(config_raw, config_type="jupyter")
base_dir = Path.cwd()
output_base_dir = Path(base_dir, "output")
wo_hydra_dir = Path(output_base_dir, "without_hydra")
out_dir_final = create_output_directory(wo_hydra_dir, "S", "2", "0.01", "47", "1000", "example")
Preprocessing#
[3]:
# preprocess the data
preprocess_data, userinput_data = PreProcess.get_variables(config_raw, out_dir_final)
Preprocessing...
User Configuration Data Setup Complete...
Geometry Data Setup Complete...
Results Data Setup Complete...
Time step set to: 47s
Applied Initial & Boundary Conditions...
Preprocessing done.
Model settings#
for more info on all the model parameters have a look at the documentation: userinput
[4]:
from spyice.parameters.user_input import Constants
# Choose between real and test constants
constants = Constants.REAL.value
Algae model
Algae model type : Multiple layers or Single layer
Radiation model
Only diffusion model (without advection)
Salinity Model
[5]:
# change user input data
userinput_data.is_diffusiononly_equation = True
userinput_data.is_algae_equation = False
userinput_data.is_radiation_equation = False
userinput_data.is_salinity_equation = False
userinput_data.algae_model_BAL_type = "all" # or "all" or "single"
[6]:
# --- Initial Conditions ---
from spyice.parameters.user_input import InitialTemperature, InitialSalinity, InitialLiquidFraction
userinput_data.initial_temperature = InitialTemperature.T_S.value
userinput_data.initial_salinity = InitialSalinity.S34.value
userinput_data.initial_liquid_fraction = InitialLiquidFraction.P1.value
[7]:
from spyice.parameters.user_input import BoundaryConditionType
from spyice.parameters.user_input import TopTemperatureType
userinput_data.boundary_condition_type = BoundaryConditionType.DIRICHLET.value
userinput_data.temperature_top_type = TopTemperatureType.STEFAN.value
userinput_data.boundary_top_temperature = 265
[8]:
from spyice.parameters.user_input import LiquidusRelation
userinput_data.liquidus_relation_type = LiquidusRelation.NORMAL.value
[9]:
preprocess_data = PreProcess.update_preprocess_dataclass(preprocess_data, userinput_data)
Time step set to: 47s
Updated Initial & Boundary Conditions...
Run model#
[10]:
# run the model
results_data = SeaIceModel.get_results(preprocess_data, userinput_data)
analysis_data = Analysis.get_error_results(
t_k_diff=results_data.t_k_diff,
t_stefan_diff=results_data.t_stefan_diff,
residual=results_data.residual_voller_all,
temperature_mushy=results_data.t_k_iter_all,
phi_mushy=results_data.all_phi_iter_all,
salinity_mushy=results_data.s_k_iter_all,
output_dir=out_dir_final,
)
Model run complete and Ready for Analysis.
Residuals exported successfully.
Visualization of Model:#
for more info on other visualization options look at: visualize model
[11]:
model_visualization_object = VisualiseModel(
user_input_dataclass=userinput_data,
results_dataclass=results_data,
error_analysis_dataclass=analysis_data,
)
Visualisation object created...
[12]:
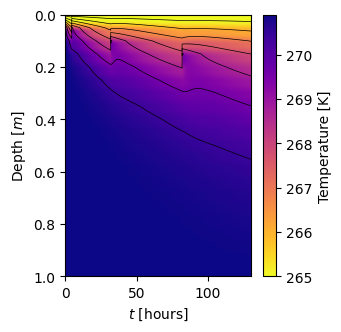
model_visualization_object.plot_temperature_heatmap(savefig=False, show=True)
Plotting Temperature heatmap...
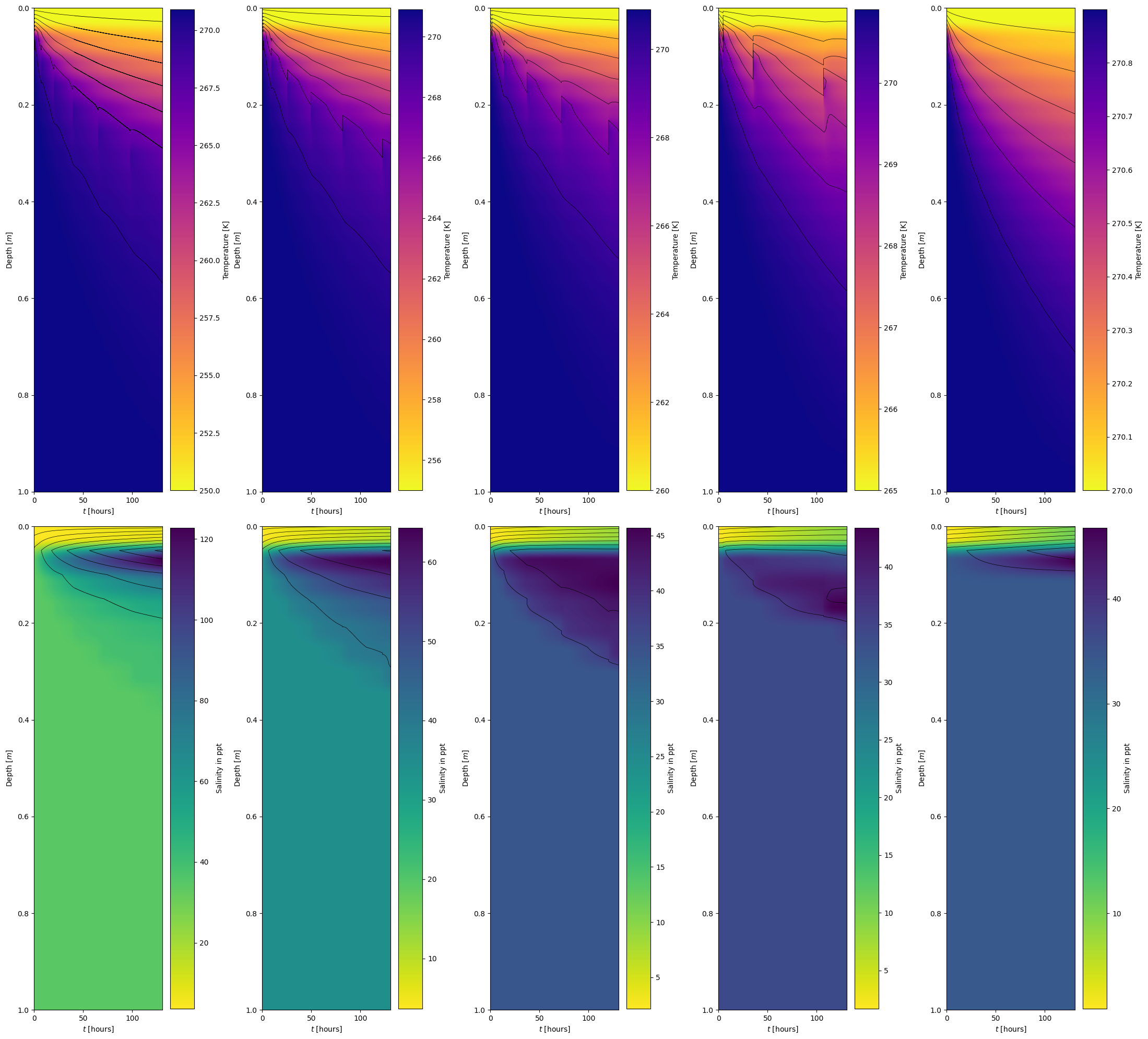
Comparing sea ice model for different temperatures#
[13]:
boundary_temperatures = [250.0, 255.0, 260.0, 265.0, 270.0]
fig, ax = plt.subplots(2, 5, figsize=(22,20))
for i, temp_bc in enumerate(boundary_temperatures):
preprocess_data, userinput_data = PreProcess.get_variables(config_raw, out_dir_final)
userinput_data.boundary_top_temperature = temp_bc
preprocess_data = PreProcess.update_preprocess_dataclass(preprocess_data, userinput_data)
results_data = SeaIceModel.get_results(preprocess_data, userinput_data)
analysis_data = Analysis.get_error_results(
t_k_diff=results_data.t_k_diff,
t_stefan_diff=results_data.t_stefan_diff,
residual=results_data.residual_voller_all,
temperature_mushy=results_data.t_k_iter_all,
phi_mushy=results_data.all_phi_iter_all,
salinity_mushy=results_data.s_k_iter_all,
output_dir=out_dir_final,
)
model_visualization_object = VisualiseModel(
user_input_dataclass=userinput_data,
results_dataclass=results_data,
error_analysis_dataclass=analysis_data
)
model_visualization_object.plot_temperature_heatmap(savefig=False, show=False, ax=ax[0,i], fig=fig)
model_visualization_object.plot_salinity_heatmap(savefig=False, show=False, ax=ax[1,i], fig=fig)
plt.tight_layout()
plt.show()
Preprocessing...
User Configuration Data Setup Complete...
Geometry Data Setup Complete...
Results Data Setup Complete...
Time step set to: 47s
Applied Initial & Boundary Conditions...
Preprocessing done.
Time step set to: 47s
Updated Initial & Boundary Conditions...
Model run complete and Ready for Analysis.
Residuals exported successfully.
Visualisation object created...
Plotting Temperature heatmap...
Plotting Salinity heatmap...
Preprocessing...
User Configuration Data Setup Complete...
Geometry Data Setup Complete...
Results Data Setup Complete...
Time step set to: 47s
Applied Initial & Boundary Conditions...
Preprocessing done.
Time step set to: 47s
Updated Initial & Boundary Conditions...
Model run complete and Ready for Analysis.
Residuals exported successfully.
Visualisation object created...
Plotting Temperature heatmap...
Plotting Salinity heatmap...
Preprocessing...
User Configuration Data Setup Complete...
Geometry Data Setup Complete...
Results Data Setup Complete...
Time step set to: 47s
Applied Initial & Boundary Conditions...
Preprocessing done.
Time step set to: 47s
Updated Initial & Boundary Conditions...
Model run complete and Ready for Analysis.
Residuals exported successfully.
Visualisation object created...
Plotting Temperature heatmap...
Plotting Salinity heatmap...
Preprocessing...
User Configuration Data Setup Complete...
Geometry Data Setup Complete...
Results Data Setup Complete...
Time step set to: 47s
Applied Initial & Boundary Conditions...
Preprocessing done.
Time step set to: 47s
Updated Initial & Boundary Conditions...
Model run complete and Ready for Analysis.
Residuals exported successfully.
Visualisation object created...
Plotting Temperature heatmap...
Plotting Salinity heatmap...
Preprocessing...
User Configuration Data Setup Complete...
Geometry Data Setup Complete...
Results Data Setup Complete...
Time step set to: 47s
Applied Initial & Boundary Conditions...
Preprocessing done.
Time step set to: 47s
Updated Initial & Boundary Conditions...
Model run complete and Ready for Analysis.
Residuals exported successfully.
Visualisation object created...
Plotting Temperature heatmap...
Plotting Salinity heatmap...
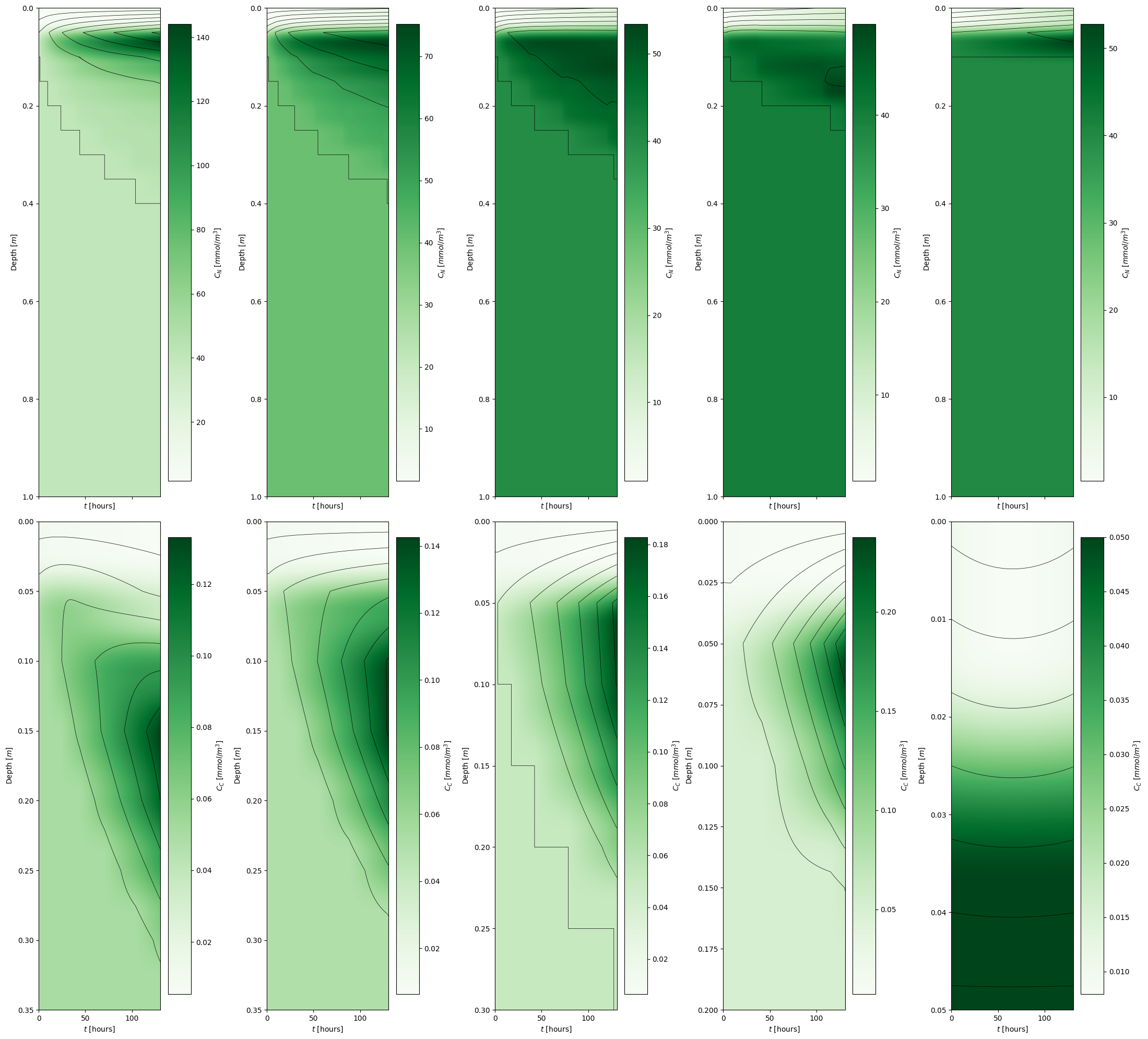
Effect of temperature on microalgae growth#
[14]:
boundary_temperatures = [250.0, 255.0, 260.0, 265.0, 270.0]
fig, ax = plt.subplots(2, 5, figsize=(22,20), sharex=True)
for i, temp_bc in enumerate(boundary_temperatures):
preprocess_data, userinput_data = PreProcess.get_variables(config_raw, out_dir_final)
userinput_data.boundary_top_temperature = temp_bc
preprocess_data = PreProcess.update_preprocess_dataclass(preprocess_data, userinput_data)
results_data = SeaIceModel.get_results(preprocess_data, userinput_data)
analysis_data = Analysis.get_error_results(
t_k_diff=results_data.t_k_diff,
t_stefan_diff=results_data.t_stefan_diff,
residual=results_data.residual_voller_all,
temperature_mushy=results_data.t_k_iter_all,
phi_mushy=results_data.all_phi_iter_all,
salinity_mushy=results_data.s_k_iter_all,
output_dir=out_dir_final,
)
model_visualization_object = VisualiseModel(
user_input_dataclass=userinput_data,
results_dataclass=results_data,
error_analysis_dataclass=analysis_data
)
model_visualization_object.plot_nutrient_concentration_multiplelayers(savefig=False, show=False, ax=ax[0,i], fig=fig)
model_visualization_object.plot_carbon_concentration_multiplelayers(savefig=False, show=False, ax=ax[1,i], fig=fig)
plt.tight_layout()
plt.show()
Preprocessing...
User Configuration Data Setup Complete...
Geometry Data Setup Complete...
Results Data Setup Complete...
Time step set to: 47s
Applied Initial & Boundary Conditions...
Preprocessing done.
Time step set to: 47s
Updated Initial & Boundary Conditions...
Model run complete and Ready for Analysis.
Residuals exported successfully.
Visualisation object created...
Plotting nutrient concentration of all layers: heatmap...
Plotting carbon concentration multiple layers...
Preprocessing...
User Configuration Data Setup Complete...
Geometry Data Setup Complete...
Results Data Setup Complete...
Time step set to: 47s
Applied Initial & Boundary Conditions...
Preprocessing done.
Time step set to: 47s
Updated Initial & Boundary Conditions...
Model run complete and Ready for Analysis.
Residuals exported successfully.
Visualisation object created...
Plotting nutrient concentration of all layers: heatmap...
Plotting carbon concentration multiple layers...
Preprocessing...
User Configuration Data Setup Complete...
Geometry Data Setup Complete...
Results Data Setup Complete...
Time step set to: 47s
Applied Initial & Boundary Conditions...
Preprocessing done.
Time step set to: 47s
Updated Initial & Boundary Conditions...
Model run complete and Ready for Analysis.
Residuals exported successfully.
Visualisation object created...
Plotting nutrient concentration of all layers: heatmap...
Plotting carbon concentration multiple layers...
Preprocessing...
User Configuration Data Setup Complete...
Geometry Data Setup Complete...
Results Data Setup Complete...
Time step set to: 47s
Applied Initial & Boundary Conditions...
Preprocessing done.
Time step set to: 47s
Updated Initial & Boundary Conditions...
Model run complete and Ready for Analysis.
Residuals exported successfully.
Visualisation object created...
Plotting nutrient concentration of all layers: heatmap...
Plotting carbon concentration multiple layers...
Preprocessing...
User Configuration Data Setup Complete...
Geometry Data Setup Complete...
Results Data Setup Complete...
Time step set to: 47s
Applied Initial & Boundary Conditions...
Preprocessing done.
Time step set to: 47s
Updated Initial & Boundary Conditions...
Model run complete and Ready for Analysis.
Residuals exported successfully.
Visualisation object created...
Plotting nutrient concentration of all layers: heatmap...
Plotting carbon concentration multiple layers...
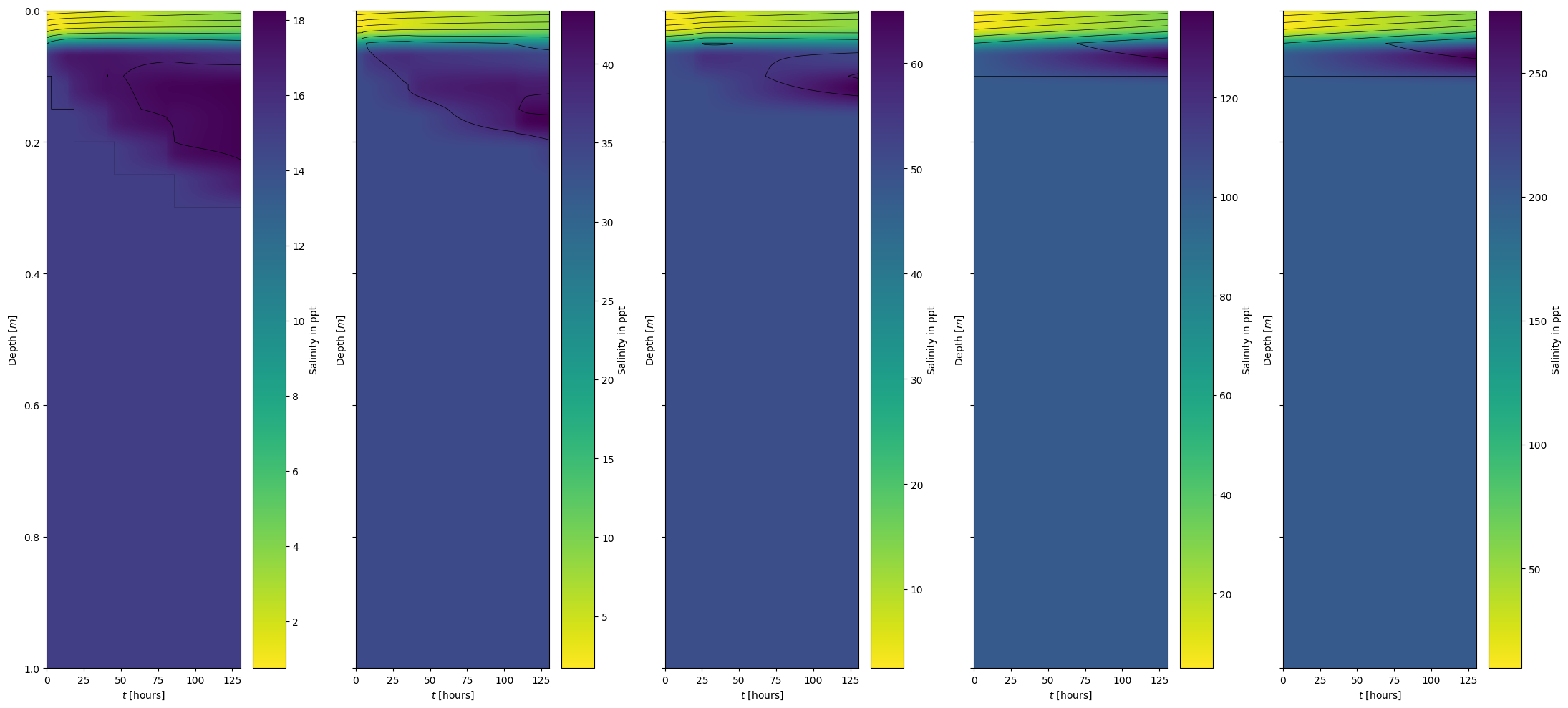
[15]:
initial_salinity = ["S15", "S34", "S50", "S100", "S200"]
boundary_salinity = [15, 34, 50, 100, 200]
fig, ax = plt.subplots(1, 5, sharey=True, figsize=(22,10))
for i, salt_ic in enumerate(initial_salinity):
preprocess_data, userinput_data = PreProcess.get_variables(config_raw, out_dir_final)
userinput_data.initial_salinity = salt_ic
userinput_data.boundary_salinity = boundary_salinity[i]
preprocess_data = PreProcess.update_preprocess_dataclass(preprocess_data, userinput_data)
results_data = SeaIceModel.get_results(preprocess_data, userinput_data)
analysis_data = Analysis.get_error_results(
t_k_diff=results_data.t_k_diff,
t_stefan_diff=results_data.t_stefan_diff,
residual=results_data.residual_voller_all,
temperature_mushy=results_data.t_k_iter_all,
phi_mushy=results_data.all_phi_iter_all,
salinity_mushy=results_data.s_k_iter_all,
output_dir=out_dir_final,
)
model_visualization_object = VisualiseModel(
user_input_dataclass=userinput_data,
results_dataclass=results_data,
error_analysis_dataclass=analysis_data
)
model_visualization_object.plot_salinity_heatmap(savefig=False, show=False, ax=ax[i], fig=fig)
plt.tight_layout()
plt.show()
Preprocessing...
User Configuration Data Setup Complete...
Geometry Data Setup Complete...
Results Data Setup Complete...
Time step set to: 47s
Applied Initial & Boundary Conditions...
Preprocessing done.
Time step set to: 47s
Updated Initial & Boundary Conditions...
Model run complete and Ready for Analysis.
Residuals exported successfully.
Visualisation object created...
Plotting Salinity heatmap...
Preprocessing...
User Configuration Data Setup Complete...
Geometry Data Setup Complete...
Results Data Setup Complete...
Time step set to: 47s
Applied Initial & Boundary Conditions...
Preprocessing done.
Time step set to: 47s
Updated Initial & Boundary Conditions...
Model run complete and Ready for Analysis.
Residuals exported successfully.
Visualisation object created...
Plotting Salinity heatmap...
Preprocessing...
User Configuration Data Setup Complete...
Geometry Data Setup Complete...
Results Data Setup Complete...
Time step set to: 47s
Applied Initial & Boundary Conditions...
Preprocessing done.
Time step set to: 47s
Updated Initial & Boundary Conditions...
Model run complete and Ready for Analysis.
Residuals exported successfully.
Visualisation object created...
Plotting Salinity heatmap...
Preprocessing...
User Configuration Data Setup Complete...
Geometry Data Setup Complete...
Results Data Setup Complete...
Time step set to: 47s
Applied Initial & Boundary Conditions...
Preprocessing done.
Time step set to: 47s
Updated Initial & Boundary Conditions...
Model run complete and Ready for Analysis.
Residuals exported successfully.
Visualisation object created...
Plotting Salinity heatmap...
Preprocessing...
User Configuration Data Setup Complete...
Geometry Data Setup Complete...
Results Data Setup Complete...
Time step set to: 47s
Applied Initial & Boundary Conditions...
Preprocessing done.
Time step set to: 47s
Updated Initial & Boundary Conditions...
Model run complete and Ready for Analysis.
Residuals exported successfully.
Visualisation object created...
Plotting Salinity heatmap...